Hi,
I’m trying to read several files from the internet via the File Reader node in a loop, where the URL to the file is controlled by a variable. It works fine in principle, but stops for one file, telling me the file does not exist. This is the file where it fails: https://www-metrabase.ch.cam.ac.uk/metrabaseui/pageview/compounds_by_target/search_results.txt?protein_name=PEPT1&action_type=substrate
If I try to read this file individually, I first get the same error on the configuration page of the File Reader, but when I increase the Timeout it shows the content in the preview window and reads the file when I execute the node.
I tried increasing the timeout first, then enable the variable, but if I do this it doesn’t remember the timeout settings. As alternative, I also tried to download the files first with the Transfer Files node, but I have the same issue there.
Do you have any ideas how I could increase the timeout, or download several files in a loop, other than manually downloading them first or adding a File Reader node for each individual file?
Thanks,
Daniela
Hello,
Have you, by any chance, tried Chunk Loops and the Try-Catch nodes?
Let me know if this works for your use case.
Hi @victor_palacios,
I’m using a table row to variable loop start, as the file reader does not take a table as input. The try/catch will only help insofar, that the workflow doesn’t break. But I know that there is data available with this link, I just cannot retrieve it because the timeout is not long enough.
I’ve created a minimum example with two links, showing that it works with the individual query, but not while using it in a loop.
Minimum example.knwf (20.3 KB)
The webpage retriever does not work as this is not a webpage, so the result is empty there. What I would need is a node to retrieve a file from a link (as the file reader can do), but with the option to set a timeout (as with the GET Request or Webpage Retriever). I think this worked with the old file reader, at least I never encountered such an issue there.
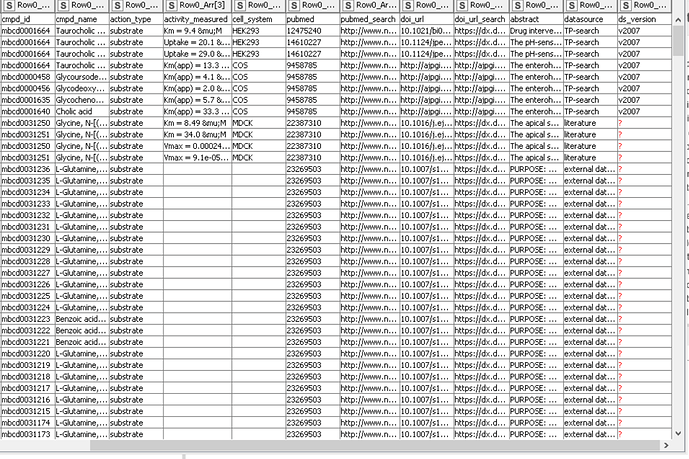
Hi @daniela_digles , is this the normal output for the extracted table, where you’ll see empty/null cells?
Hi @badger101, thank you for looking into it!
Yes, that’s ok. There are some values missing in this file, it seems there is no version information of the dataset, and no reported activity values. It’s the same if I look at the online version Metrabase | MetrabaseUI | Search results.
@daniela_digles While I could replicate the issue, I don’t know why the second URL file can’t be read with the File Reader (nor can it be read with other reader nodes I’ve tried).
Having said that, until someone can look it up for you, here’s a temporary solution to download and read the files for both URLs. Considering the files follow a .txt format, I utilized a node from the Vernalis extension. If you haven’t downloaded the extension yet, you’ll be prompted to download & install it once you tried to open this workflow I’m providing:
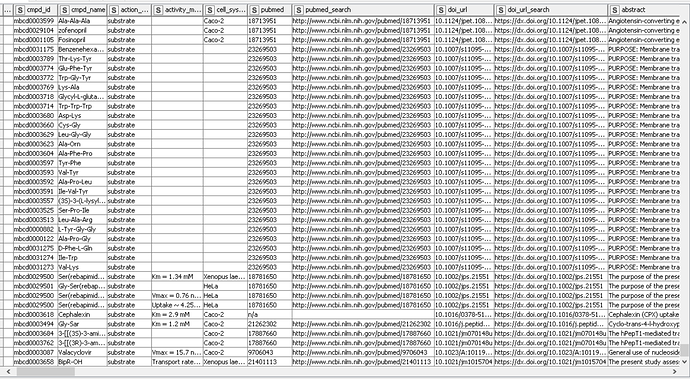
Here’s how the final table looks like (a snapshot):
Hope it helps while you wait for other solutions!
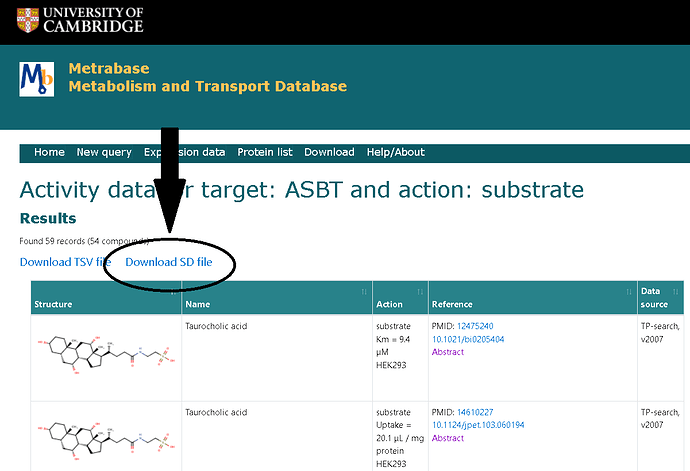
Extending to what I’ve written above, you can also try the URL links for the SD files instead of the TSVs from the Cambridge webpage.
Knime should be able to read them via this node, although I haven’t tested this alternative :
Thank you! I wasn’t aware of this node. With this node I can download all my files and extract the needed data for now. Still, it looks quite complicated to get to the needed format, so I hope that at some point the file reader node (and also related nodes such as the Transfer Files node) will work with an increased time out. @victor_palacios, can you please add this as a bug report or feature request?
Indeed, I actually need both files here, as the TSV file contains the data, and the SD file the structure. The SDF Reader node fails as well with the larger file. However, knowing now that Vernalis nodes don’t mind this, I found out that the Load SD-Files node works fine.
This topic was automatically closed 90 days after the last reply. New replies are no longer allowed.