Greetings KNIME community,
I have (I hope) a relatively simple problem, but I can’t seem to figure out how to get around it…
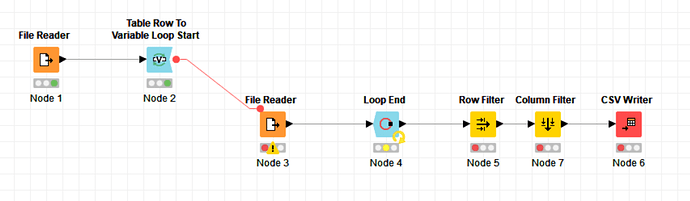
I am a chemist working on big data-related project. The ZINC database provides a useful cache of molecules to use with molecular modeling and other related tasks. You can download these for free, but the download process is a bit laborious. One must download a large number of “tranches”, each containing a few hundred or a few thousand compounds. I would like to automate this process so I constructed a simple workflow to accomplish this:
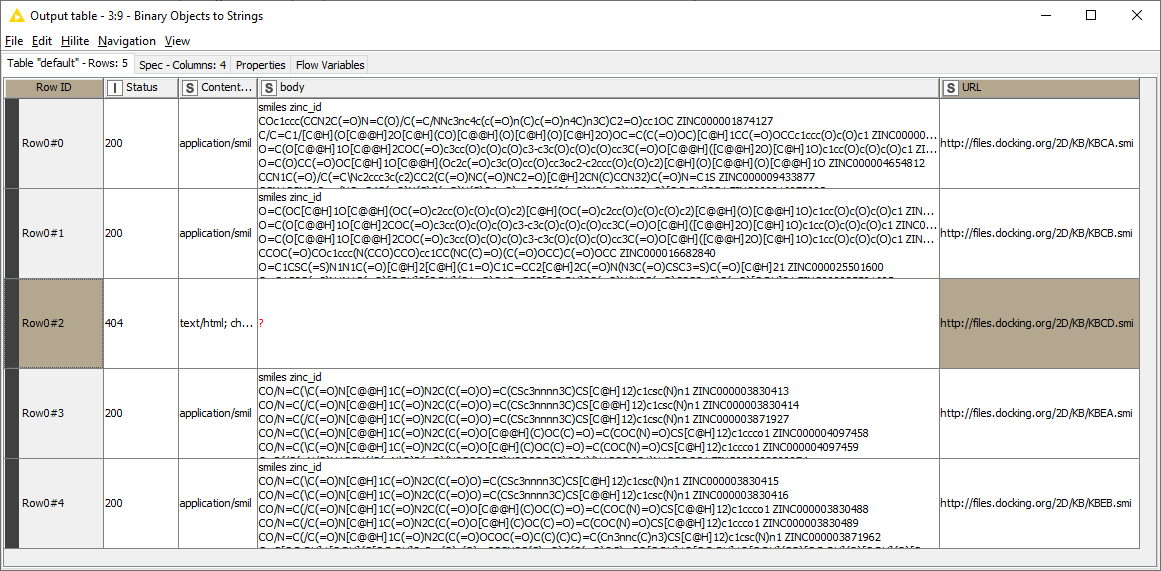
The first file reader takes the list of URL addresses generated by the ZINC database and passes them to the variable loop. The loop contains its own reader that then uses the URL list to read the data and compile it into one list. The filters just get rid of some unwanted formatting; the writer then creates the file I want. It works great until I run into one particular address that is not valid. Then the whole workflow stops. I get the following warning generated by the reader in the loop: “Execute failed: Not a file or knime URL: 'http://files.docking.org/2D/KB/KBCA.smi”. I tried accessing that file with my browser, and it looks like the address is simply corrupt.
I tried a couple work-arounds using try/catch loops and using the File Meta Info node to pre-filter the list of URLs, but neither was successful. The try/catch approach gets stuck, I think, because the warning generated isn’t actually an error, just a warning, And the meta data approach doesn’t work because the “exists” column does not populate for remote sites.
Can anybody think of a good way I can have my loop continue if it comes upon a corrupt address? I don’t mind skipping the data, but it is quite inconvenient if I need to keep going back to trim out the corrupt locations from my original list of URLs.
Thanks for your help!
-JW