Hi,
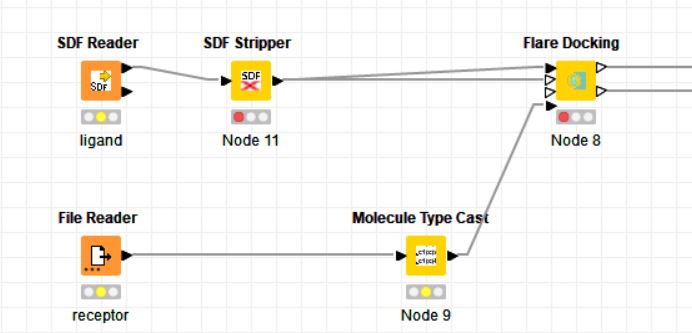
I´m using Flare Docking node (Cresset) to perform some dockings. However, it halts with the following message:
ERROR Flare Docking 4:6 Execute failed: Error (exit code 255) : Error (exit code 255) running
Error (exit code 255) running
Error: module ‘rdkit.Chem’ has no attribute ‘Mol’
---- Exception Information For Developers ----
Traceback (most recent call last):
File “/apps/cresset/Flare/bin/…/pyflare-scripts/docking.py”, line 68, in main
project, ligands_to_dock_role = _configure_process(docking, args)
File “/apps/cresset/Flare/bin/…/pyflare-scripts/docking.py”, line 379, in _configure_process
role.ligands.extend(flare.read_file(args.reference, _find_format(args.reference, args)))
AttributeError: module ‘rdkit.Chem’ has no attribute ‘Mol’
I got Knime 4.7.5 and RDKit 2022.09.3. Linux LT 20.04 as OS
Any ideas of what can cause this situation and how to sort it?
Thanks a lot,
E77