@toscanomatheus this might be an interesting case. I will have to have a closer look at your specific file.
I startet to look into the question pof the Python Labs nodes and Date and Time variables (again) and indeed there seems to be a problem with the handling of types. In a previous discussion I toyed around with several formats and then exported the data thru Parquet files and brought the data back to KNIME which worked (mostly):
This time I tried two things with string formats exported from KNIME to Python and there transformed them into Timestamps with either Pandas or PyArrow. Again this worked within Python and you could export that with the help of Parquet but the date time formats would fail when you wanted to bring them back to KNIME.
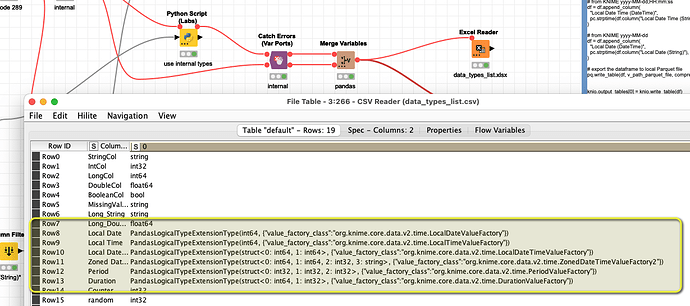
The Date and Time variables brought from KNIME to Python do have some very specific formats and I am not sure they can be used in Python in a good way:
What does work is to convert specific strings to Python readable Formats. Here shown in a Jupiter notebook but then within the Python KNIME node:
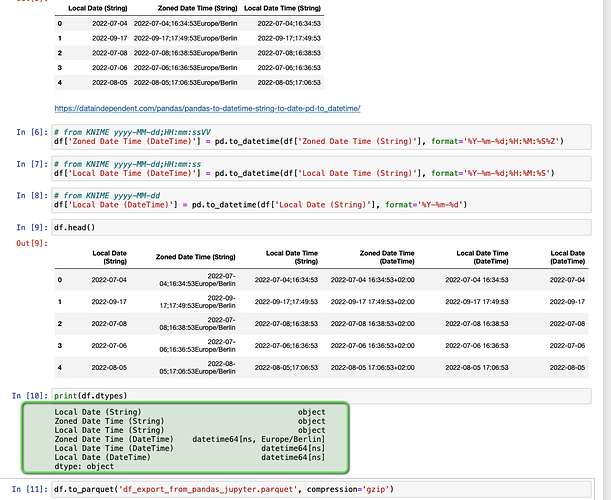
# using Pandas
# from KNIME yyyy-MM-dd;HH:mm:ss
df['Local Date Time (DateTime)'] = pd.to_datetime(df['Local Date Time (String)'], format='%Y-%m-%d;%H:%M:%S')
The resulting “datetime64[ns]” or “datetime64[ns, Europe/Berlin]” format seems not to be supported for conversion back to KNIME, though it can be stored as parquet.
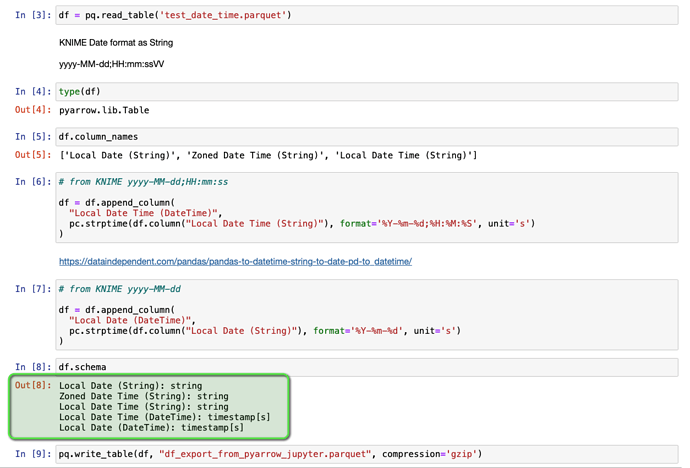
The same is true if you employ PyArrow to convert the strings to Date and Time:
# using PyArrow
# from KNIME yyyy-MM-dd;HH:mm:ss
df = df.append_column(
"Local Date Time (DateTime)",
pc.strptime(df.column("Local Date Time (String)"), format='%Y-%m-%d;%H:%M:%S', unit='s')
)
@carstenhaubold I think the handling of date and time variables between KNIME and Python might have to be improved. I think standard timestamps from Pandas and PyArrow should be supported so you could bring back results in such a format. Otherwise you would have to resort to parquet or string variables.
in the sub-folder /data/ there are two Jupyter notebooks to try a few things with date and time variables with
Pandas: knime_py_pandas_date_time_columns.ipynb
PyArrow: knime_py_pyarrow_date_time_columns.ipynb