Hi everyone,
I have the following task at hand.
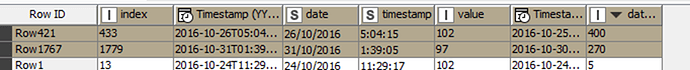
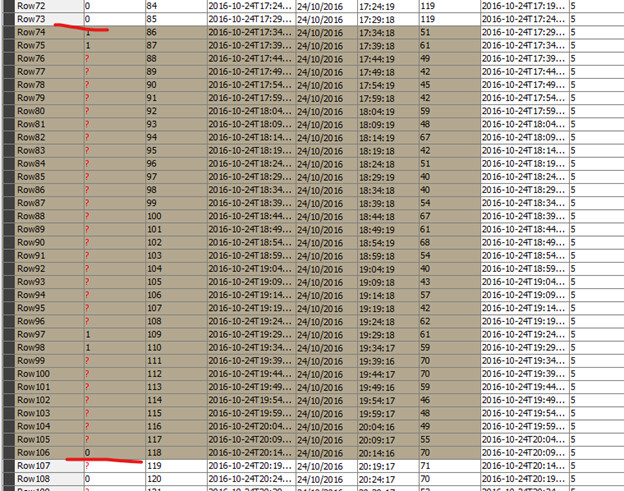
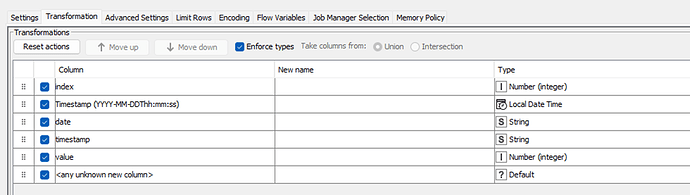
I have a dataset consisting of the following columns: obs_index, datetime, date, time, value.
What I want to achieve: I need to ‘event flag’ (see GOAL column in the below screenshots) based on the value and duration conditions, using only KNIME nodes (not R or Python snippets) if possible.
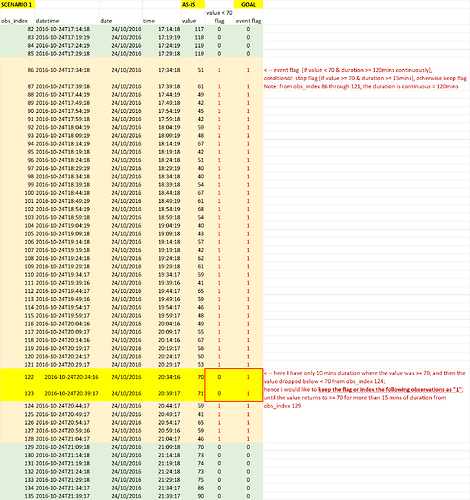
The logic needed to be implemented is as follows: flag the event [if value < 70 & duration >= 120 mins continuously], stop flagging [if value >=70 & duration >= 15mins continuously], otherwise keep flagging.
I have prepared two scenarios for your information. Please see the below screenshots.
Scenario 1:
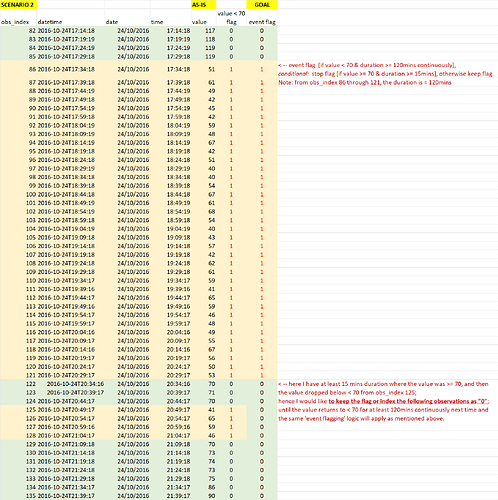
Scenario 2:
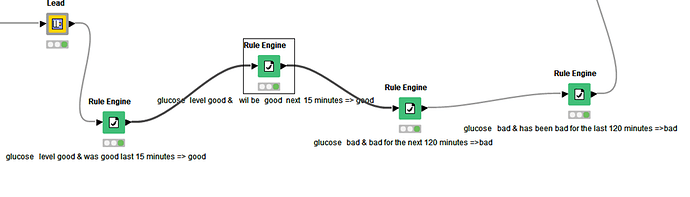
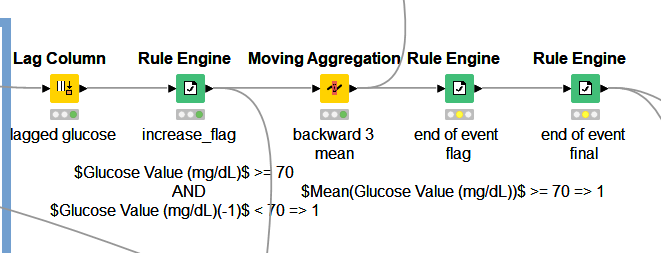
Tried method: I tried to solve the problem using the lag column, rule engine, and moving aggregation nodes (with window = 3, [mean(value)] backward window type method). Then put the condition of flagging the “end of event” row(s) wherever the value was increasing above >= 70 AND at the same time the mean(value) for the last three observations were >= 70. However, this didn’t work for all the possible >=120 observations. The workflow that was unsuccessful for me:
Let me know if you have any questions or need any clarification.
Your help is greatly appreciated! ![]()
Thank you!
Odko