Hi everyone,
I’m trying to reproduce in KNIME exactly what I do in R with:
km_res <- kmeans(data_scaled, centers = 3, nstart = 1000, iter.max = 200)
That is, run K-Means with 1000 random starts (nstart = 1000), pick the best solution (minimum total within-cluster sum of squares), and then use that model.
So far I’ve attempted:
KNIME’s built-in K-Means node
I can set k = 3, max iterations = 200, and a fixed seed, but there is no nstart or “number of restarts” parameter, nor any checkbox to output the total within-cluster sum of squares for each run.
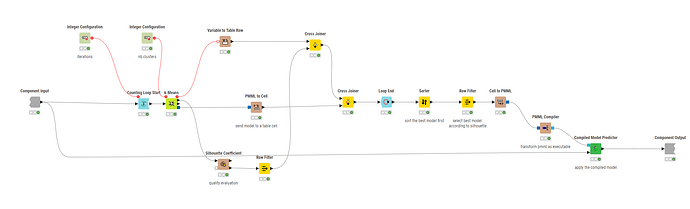
Counting Loop Start → K-Means → Variable Loop End
I tried mapping the loop index (iteration) as the seed, and hoped to collect the WSS via the K-Means node’s flow variables—but there isn’t actually an option in the K-Means node to append totalWithinSS (or similar) into flow variables.
Without that, I can’t sort/filter to pick the best run automatically.
Question:
Is there a more direct or built-in way to perform multiple random starts with KNIME’s native K-Means (or a common extension), analogous to R’s nstart? (without using Python/R snippets)
Thanks in advance for any pointers!