Hi everyone,
What I want to do:
Read data from my hard drive which contains batch data (in numpy .npz format) and output a numpy array which I want to input into a neural network executor node. The network is a very sophisticated named entity recognition deep learning model, composed of a BILSTM and a CNN (the model works perfectly fine in plane python and produces state of the art results).
What I did:
I used the Python Source node with the following code (code chunk I):
import pandas as pd
import numpy as np
path = ‘…/test_batches_casefeat_punct_unk.npz’
test_batches = np.load(path, encoding=‘latin1’)[‘arr_0’]
#output_table = test_batches
#print(output_table[0])
output_table = pd.DataFrame(data = test_batches)
print(output_table.iloc)
My Problem(s) with this:
- Like described, I need a numpy array as output, not pandas data frame
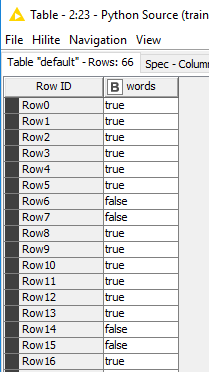
- Second: If I read the output_table from the above code with a Python Script (1 => 1) node and investigate my four colums, I only see booleans (but there should be integers), e.g. (code chunk 2):
a = input_table[‘0’]
print(a)
yields:
Row0 True
Row1 True
…
Row64 True
Row65 True
Name: 0, Length: 66, dtype: bool
but it should look like this (as a PandasDataframe):
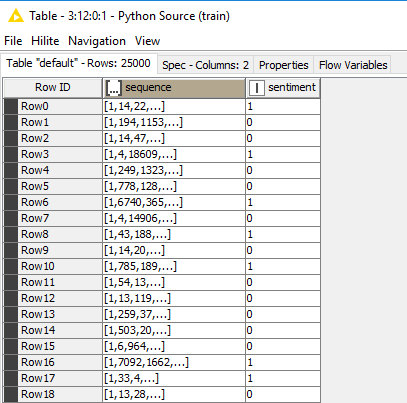
0 [[112], [2790], [3495], [7564], [549498], [217…
1 [[[8, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0…
2 [[4], [3], [3], [3], [2], [3], [4], [3], [3], …
3 [[[4]], [[4]], [[4]], [[6]], [[2]], [[6]], [[4…
Name: 0, dtype: object
(this is the result from the print call inside of the Python Source node above in code chunk I)
So how do I get an numpy array as a node output?
And why does the Python Source node creates some weird boolean output?
Thanks in advance,
Niklas