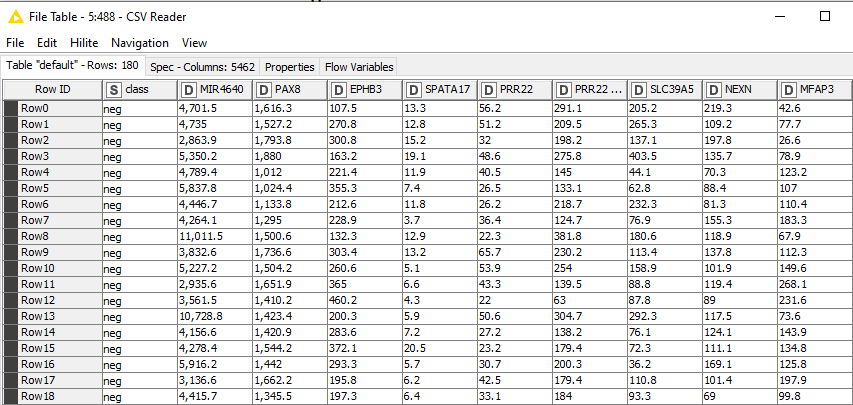
Hello, I am working with a gene expression dataset. In the image below, you can see the structure of my dataset. I am attempting to straightforwardly use the Limma package on my dataset. I am using a CSV reader to read my file, and then I am using an R snippet with the following code to run the Limma package. However, I am encountering different errors, with the error below being the most persistent. Should I be performing more data transformation steps before feeding my data into the R snippet with the limma package? If someone could tell me where I am making a mistake and how I can fix it, it would be great. Or if someone already has an existing limma node for knime for Windows that would be great as well.
code
"
Convert the ‘class’ column to a factor
kIn$class ← factor(kIn$class)
Create a design matrix
design ← model.matrix(~0 + kIn$class)
Extract the expression data
exprs_data ← as.matrix(kIn[, -1]) # Convert to a matrix
Load the limma package
if (!requireNamespace(“BiocManager”, quietly = TRUE))
install.packages(“BiocManager”)
BiocManager::install(“limma”)
library(limma)
Fit the linear model
fit ← lmFit(exprs_data, design)
Apply empirical Bayes moderation
fit ← eBayes(fit)
Extract the results
results ← topTable(fit, coef=“kIn$classpos”, number=nrow(exprs_data))"
This is the error
“ERROR R Snippet 5:489 Execute failed: Failed to evaluate the script:
Error : Error in lmFit(exprs_data, design) :
row dimension of design doesn’t match column dimension of data object”