Hi Evert,
Thanks for bringing this subject to the KNIME forum which sounds very interesting for the field of cheminformatics.
I had a look at the links you mentioned and as far as I see, the calls to this Python library are done based on command line calls (as stated in the ChemProp Tutorial Data section). In this case you are not obliged to call it through a Python script but any node allowing command-line execution would do the job.
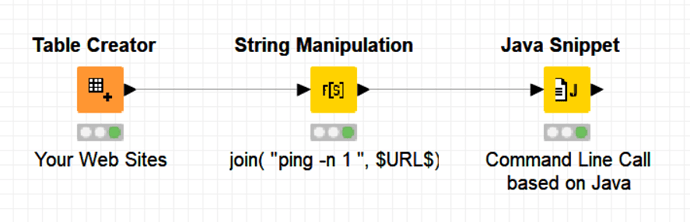
Recently, I posted a solution based on Java (and hence with no dependency on Python) to do command-line calls which can recover the output terminal results generated by the command line:
(It was used to ping URLs but can be used for any other command-line use as i.e. it is required in your case)
Nevertheless, in your particular case I see that you would need first to save the table you want to pass as a CSV file and secondly to read the results as a CSV file too once your command line has finished its execution.
The solution I implemented waits until the end of the execution and tells you in the end if everything went ok with command-line outputs and errors if any as well. It could be an option for your need here.
Otherwise you could always do the same using a Python Script node but I do not see the advantage since what you execute is not a Python code but a command-line program. Other options are available as stated by @ScottF in the same thread.
I’ll be happy to further help if needed.
Hope this already helps
Best
Ael