Recently I posted a topic to make Knimers aware of a EDA package called Sweetviz. Here’s the GitHub URL : GitHub - fbdesignpro/sweetviz: Visualize and compare datasets, target values and associations, with one line of code. .
Here’s a sample script using the Titanic survivor data:
import sweetviz
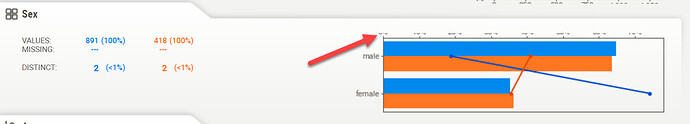
If I run this script in a Knime Python Script node, the HTML output truncates the chart headers:
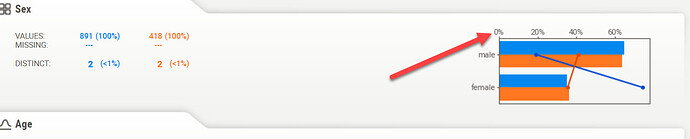
If I run the same script in Anaconda Spyder, the chart is displayed correctly. I’m using the same Python environment in both Knime and Spyder:
Any thoughts?
Hey @rfeigel ,
I ran sweetviz.analyze() on a sample dataset in Windows 10 with the script below and it rendered properly. It opened a new window in my default web browser with the rendered report. Here’s the code I used for the Python Script node:
import sweetviz
# Copy input to output
output_table_1 = input_table_1.copy()
my_report = sweetviz.analyze(output_table_1)
my_report.show_html()
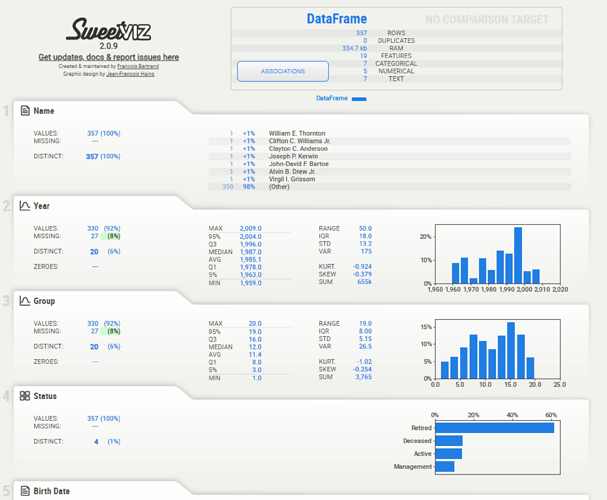
Here’s the output:
I did also see a scaling option in their documentation which reads:
scale: Use a floating-point number (scale= 0.8 or None) to scale the entire report. This is very useful to fit reports to any output.
Hope this helps!
Cheers,
@sjporter
2 Likes
I tried varying the scale and switching bewteen vertical and widescreen. Nothing helps. Still have same problem. It plots, but I still have the same truncation problem. Here’s my current workflow:
import sweetviz
output_table_1 = input_table_1.copy()
Rather than using pandas to import the file I tried using a csv reader node to feed the Python Script node.
My csv file (changed to a txt file since I can’t upload a csv file) train.txt (59.8 KB) is attached.
Hey @rfeigel ,
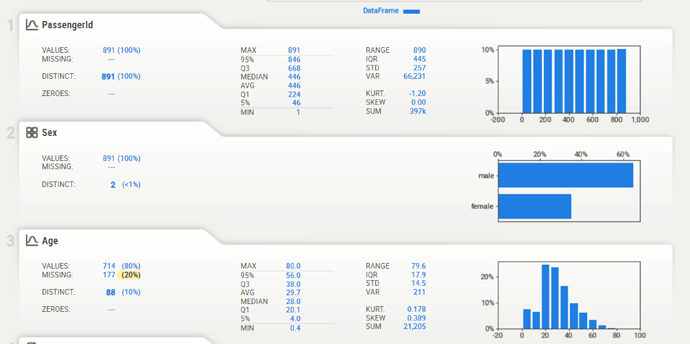
I ran your script against the same test data (OS: Windows 10, Browser: Google Chrome) and it appears to be rendering properly:
import sweetviz
output_table_1 = input_table_1.copy()
my_report = sweetviz.analyze(output_table_1)
my_report.show_html(
filepath="SWEETVIZ_REPORT.html",
open_browser=True,
layout="widescreen",
scale=1.2
)
Which OS / browser are you using?
1 Like
I’m using Windows 10 and Chrome.
I just tried MS Edge and MS Explorer and have the same problem. I’m really puzzled since it runs fine in Anaconda Spyder with the Chrome browser.
MS Edge is Chromium-based just like Google Chrome, so if you want to determine if your browser is the root cause I’d recommend trying Firefox or Internet Explorer.
If that doesn’t lead to any insights, could you please try creating a conda environment based on the definition below and use the Conda Environment Propagation node to load it? I’m using Python 3.6.12 for this environment.
name: py36_knime_sweetviz
channels:
- defaults
dependencies:
- appnope=0.1.2=py36hecd8cb5_1001
- arrow-cpp=0.11.1=py36hcacac7f_1
- attrs=20.3.0=pyhd3eb1b0_0
- backcall=0.2.0=pyhd3eb1b0_0
- blas=1.0=mkl
- bzip2=1.0.8=h1de35cc_0
- ca-certificates=2021.1.19=hecd8cb5_0
- cairo=1.14.12=hc4e6be7_4
- certifi=2020.12.5=py36hecd8cb5_0
- cycler=0.10.0=py36hecd8cb5_0
- decorator=4.4.2=pyhd3eb1b0_0
- fontconfig=2.13.1=ha9ee91d_0
- freetype=2.10.4=ha233b18_0
- gettext=0.19.8.1=hb0f4f8b_2
- gflags=2.2.2=h0a44026_0
- glib=2.66.1=h9bbe63b_0
- glog=0.3.5=h0a44026_1
- icu=58.2=h0a44026_3
- importlib-metadata=2.0.0=py_1
- importlib_metadata=2.0.0=1
- intel-openmp=2019.4=233
- ipython=7.1.1=py36h39e3cac_0
- ipython_genutils=0.2.0=pyhd3eb1b0_1
- jedi=0.13.3=py36_0
- jpeg=9b=he5867d9_2
- jsonschema=3.2.0=py_2
- jupyter_core=4.7.1=py36hecd8cb5_0
- kiwisolver=1.3.1=py36h23ab428_0
- libboost=1.67.0=hebc422b_4
- libcxx=10.0.0=1
- libedit=3.1.20191231=h1de35cc_1
- libevent=2.1.8=hddc9c9b_1
- libffi=3.3=hb1e8313_2
- libgfortran=3.0.1=h93005f0_2
- libiconv=1.16=h1de35cc_0
- libpng=1.6.37=ha441bb4_0
- libtiff=4.1.0=hcb84e12_0
- libxml2=2.9.10=h7cdb67c_3
- lz4-c=1.8.1.2=h1de35cc_0
- mkl=2019.4=233
- mkl-service=2.3.0=py36h9ed2024_0
- mkl_fft=1.2.0=py36hc64f4ea_0
- mkl_random=1.1.1=py36h959d312_0
- nbformat=4.4.0=py36_0
- ncurses=6.2=h0a44026_1
- olefile=0.46=py36_0
- openssl=1.1.1j=h9ed2024_0
- parso=0.8.1=pyhd3eb1b0_0
- pcre=8.44=hb1e8313_0
- pexpect=4.8.0=pyhd3eb1b0_3
- pickleshare=0.7.5=pyhd3eb1b0_1003
- pip=20.3.3=py36hecd8cb5_0
- pixman=0.40.0=haf1e3a3_0
- prompt_toolkit=2.0.10=py_0
- ptyprocess=0.7.0=pyhd3eb1b0_2
- pyarrow=0.11.1=py36h0a44026_0
- pygments=2.7.4=pyhd3eb1b0_0
- pyparsing=2.4.7=pyhd3eb1b0_0
- pyrsistent=0.17.3=py36haf1e3a3_0
- python=3.6.12=h26836e1_2
- python-dateutil=2.7.5=py36_0
- pytz=2021.1=pyhd3eb1b0_0
- readline=8.1=h9ed2024_0
- setuptools=52.0.0=py36hecd8cb5_0
- six=1.15.0=py36hecd8cb5_0
- snappy=1.1.8=hb1e8313_0
- sqlite=3.33.0=hffcf06c_0
- statsmodels=0.11.1=py36haf1e3a3_0
- thrift-cpp=0.11.0=hd79cdb6_3
- tk=8.6.10=hb0a8c7a_0
- tornado=6.1=py36h9ed2024_0
- traitlets=4.3.3=py36_0
- wcwidth=0.2.5=py_0
- wheel=0.36.2=pyhd3eb1b0_0
- xz=5.2.5=h1de35cc_0
- zipp=3.4.0=pyhd3eb1b0_0
- zlib=1.2.11=h1de35cc_3
- zstd=1.3.7=h5bba6e5_0
- pip:
- importlib-resources==5.1.2
- jinja2==2.11.3
- markupsafe==1.1.1
- matplotlib==3.3.4
- numpy==1.19.5
- pandas==1.1.5
- patsy==0.5.1
- pillow==8.1.2
- pytesseract==0.3.7
- scipy==1.5.4
- sweetviz==2.0.9
- tqdm==4.59.0
Chrome, Edge, Internet Explorer and Firefox all don’t work. I’ll try your environment when I get time, but frankly its probably just easier to run in Anaconda which works for me.
That’s understandable. If anyone else in the community has a couple free minutes to try out the sweetviz package and see how it turns out, perhaps we could determine if this issue is specific to your computer or something larger in scope.
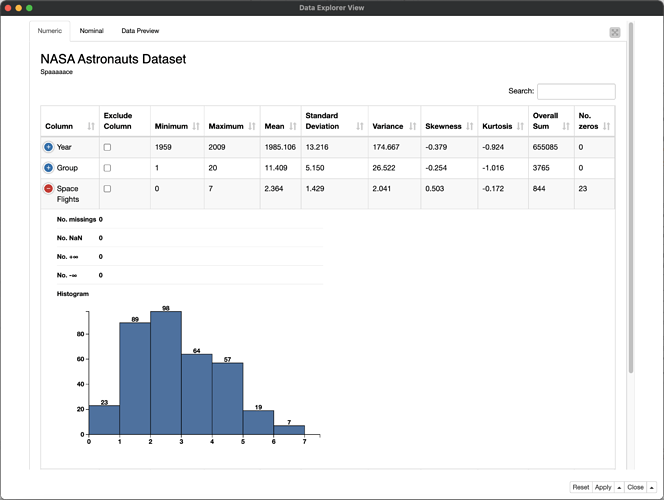
I’m sure there are specific features you want to use that the sweetviz package offers, but it’s worth mentioning that the Data Explorer node has a number of overlapping features for data profiling in case you haven’t tried it out yet:
Cheers,
@sjporter
1 Like
Thanks. The File Explorer node is very useful, but doesn’t have all the functionality of Sweetviz.
system
September 18, 2021, 6:28am
11
This topic was automatically closed 182 days after the last reply. New replies are no longer allowed.