Hello all,
I am trying to enumerate a virtual library using the RDKit Two Component Reaction node in Knime.
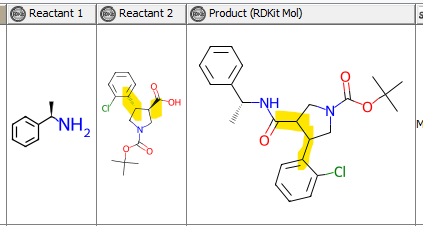
The enumeration is based on an amide coupling of a chiral amine and different acidic building blocks.
Although the chirality of some building blocks is defined, I am unable to retain this information in the enumerated products. Interestingly, the chirality of the amine is retained.
I am using Knime 5.3.3 installed automatically with the Maestro (Schrodinger).
Can you please advise?
Many thanks in advance,
Kind regards,
Silvia
Example of workflow
Enumeration_WF.knwf (19.1 KB)
Example of acidic BBs
CC(C)(C)OC(=O)N1CC@HC(=O)O
CC(=O)NC@@HC(=O)O
Amine: NC@HC1=CC=CC=C1