I am trying to create new structures from an existing structure starting from Canonical SMILES string. I am able to split the string into unique columns using Cell Character Splitter but am struggling with how to randomly modify (add column to random position from dictionary, delete random column, modify random column from dictionary). Any suggestions are appreciated such as use of random column flow variable to guide selection.
Welcome to the forum, @KAV.
What exactly do you mean when you say that you’re “trying to create new structures from an existing structure starting from Canonical SMILES string”? Do you have any examples that you can share?
I don’t quite understand all the column manipulation you mentioned having to do. At first glance, it seems to me that this kind of work would be better achieved with one of the RDKit reaction nodes.
@elsamuel A SMILES string is a character string that describers a molecule (no space). By changing one character in the string, it describes a new molecule that is different from the original molecule. I would like to do this randomly (change one character and write it out). I don’t believe RDKit has this capability as RDKit is fingerprint driven (numerical).
Molecular fingerprints directly encode molecular structure in a series of binary bits that represent the presence or absence of particular substructures in the molecule and as such do not allow single atom changes.
I am quite familiar with the various chemical representations. I’m also familiar with using reaction SMARTS to modify molecules in KNIME, which is why I mentioned it as a potential solution.
Can you provide specific examples of what you’re trying to accomplish?
Thanks for your insight, greatly appreciated. Basically I want to start with this smiles string
NC1=NC2=CC=C(Br)C=C2C(OC3CCN(C)CC3)=C1
and end up with this smiles string
NC1=NC2=CC=C(Br)C=C2C(OC3CCN(O)CC3)=C1
where one character of the string is chosen, modified from a dictionary or list or possible choices and done at random (both the selection of the atom and the picking from the dictionary).
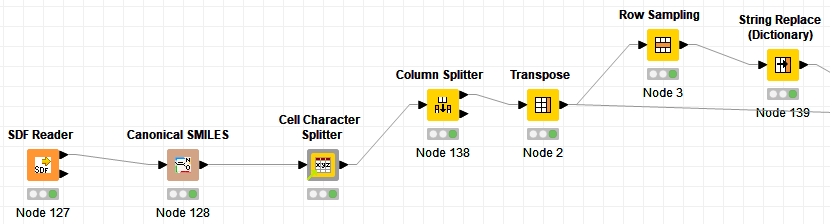
What I have right now is based on individual cell manipulation. I read in a sdf file, create a connonical smiles string, use a cell character splitter node, remove the structure and smiles columns with column splitter, transpose columns to rows (pivot), sample a random row using row sampling, then replace with string replace dictionary (this part is not random but rather sequential within my list). I would like to make the dictionary step random as well.
Thank you!
What do the dictionary entries for the string replacement look like? How many entries are there?
For now I am working with just these four lines (comes back with no change if nothing is found for example for a parenthesis character). Likely not the best way to do this as a randomization is not possible.
C, N, O, S, Cl, Br, I, F
N, C, O, S, Cl, Br, I, F
O, C, N, S, Cl, Br, I, F
S, C, N, O, Cl, Br, I, F
Would you be able to upload the workflow? That would make things much easier.
Not really an answer to your question but have you looked at molpher and molpher-lib? The latter being accessible via python (and hence also KNIME).
It might help with your ultimate goal?
Had not come across this one yet, will give it a look, thanks!
@elsamuel I am reworking the workflow to make it random in its selection of replacement. I figured out how to modify a single character and am dealing with re-insertion of a this now matched row (new character with same row and column header names in a second table).
OK, I have solved most of the problems but ran into one I did not anticipate. I can split a smiles string into characters and recombine them back into a smiles string but when I have a two character element like Cl I get two columns “C” and “l” instead of one with “Cl”. Any suggestions on how to recombine these back into a single column without losing my place in the table (select two columns, combine and replace original column left with new combined text string while deleting column right).
![]()
Hello @KAV,
have you managed to solve it?
To my knowledge there is no single node to do that. However this can be easily done on couple of ways with multiple nodes. For example String Manipulation with join() function and Replace Column option followed by Column Filter.
Br,
Ivan
Or how about using Python Script node and RDKit?
m = Chem.MolFromSmiles(‘c1ccccc1O’)
em = Chem.RWMol(m)
em.ReplaceAtom(6,Chem.Atom(9))
Chem.SanitizeMol(em)
Chem.MolToSmiles(em)
Fc1ccccc1
All you need to change above is to apply the “random” replacement. If you convert your smiles to rdkit molecule before the Python Script node the SMILES conversions aren’t needed.
Sanitize step is required to verify a valid molecule was created. if not, you can drop it and try again.
Thank you, I will try this. I have not found a better solution yet that does not involve multiple nodes.
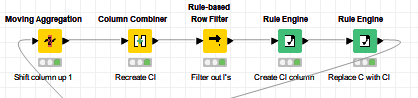
Solved the problem using only standard nodes. Basically had to use moving aggregation to create a new column shifted up one cell. Then used a column combiner to create “Cl” (and all the other combinations that don’t apply), then filtered out the original rows that contained just “l” then used two rule engines to create a column containing only the replacements (“Cl”) and replaced in the final engine.
This topic was automatically closed 182 days after the last reply. New replies are no longer allowed.