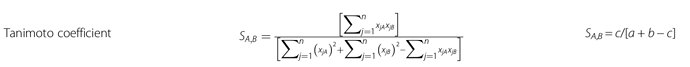Hi guys,
Is there a way in KNIME (without using java snipped nodes) to calculate Tanimoto similarity (ranging from 0 to 1) using count-based fingerprints (e.g. RDKit Morgan count-based fingerprints)?
Currently the Byte Vector Distances node doesn’t allow it.
Thanks in advance!
Gio
Hi,
There’s not a node to calculate it directly, but if you really want to avoid using a Java Snippet node (which would be the fastest and simplest), you can use some of the fingerprint operations nodes in the Vernalis extensions to calculate the Jaccard distance (essentially the Tanimoto distance) between two count vectors.
I’ve attached a workflow that does this and captures the functionality in a wrapped metanode. You can choose the columns to compare by double clicking the wrapped metanode.
count_based_tanimoto.knwf (24.0 KB)
-greg
p.s. Note that this does not handle the case that neither fingerprint has any bits set. This will lead to a NaN.
Dear Greg,
Thank you very much for your help. I didn’t know the Vernalis extensions for fingerprints operations and I’m sure they will be useful for me in the future.
To what deals with the count-based Tanimoto similarity (or distance) I would like to implement the formula attached as figure:
Here, S denotes similarities, xjA means the j-th feature of molecule A . a is the number of on bits in molecule A , b is number of on bits in molecule B , while c is the number of bits that are on in both molecules. On the left part of the figure there is the formula for continuous variables, while on the right part, the formula for dichotomous variables.
This formulas to calculate Tanimoto similarity are mentioned inter alia in the following publications:
- Willett P. J. Chem. Inf. Comput. Sci. 1998, 38, 983-996
- Bajusz D. et al. Journal of Cheminformatics (2015) 7:20
Do you know what are the advantages/disadvantages of using the formula you propose respect the one I mentioned? Could you please point me to a reference publication where the formula you provided is mentioned?
Finally, as a reference for other interested people reading this post, in order to implement the formula of Tanimoto coefficient for continuous variables (using KNIME byte vectors) reported in the attached figure (left part) and in the 2 aforementioned publications, I had to use a Java Snippet node as it seems not possible to implement it using Vernalis extensions for fingerprints operations. I don’t have much experience with Java but it seems it can be done easily. The probe and target fingerprints variables are “currentFingerprint” and “targetFingerprint”, respectively. Here it is the code:
if (currentFingerprint.length != targetFingerprint.length) {
throw new RuntimeException("Fingerprint vectors must be of the same length");
}
int n = currentFingerprint.length;
double ab = 0.0;
double a2 = 0.0;
double b2 = 0.0;
for (int i = 0; i < n; i++) {
ab += currentFingerprint[i] * targetFingerprint[i];
a2 += currentFingerprint[i] * currentFingerprint[i];
b2 += targetFingerprint[i] * targetFingerprint[i];
}
out_Tanimotosimilarity = ab / (a2 + b2 - ab);Thanks for posting this. Given that bytevector fingerprints are now over a decade old, I was confused by KNIME’s lack of Tanimoto-type similarity calcs. Your solution is a step in the right direction to solve this ongoing issue. Thank you!
A follow up question: how would you suggest converting the distance output in this workflow to a KNIME distance matrix that can be used downstream with other nodes? I’ve seen very complex solutions that involve 20 nodes of processing, writing to disk and then reading back from disk, but there has to be a better way, right?