Hi,
I get the raw data from LC-MS,such as the Groups,QC,and Blank.Now I have a question in inputting the data to the Input Files.I should choose the Groups data to the workflow or the three type data.
In the begining, I choose the example open data to excute the workflow,Now I decide to perform the workflow with the Groups(A,B,C).
Thank you!
Hi
this depends on what you want to do with the QC and blank groups. If you want to align them to e.g. remove features from Blank in the Groups, I would probably process them together.
But maybe you should start simple and just compare the groups.
What is your QC sample for, exactly?
Yes, QC should be correct the stabilization of data. But the blank and the QC shouldn’t input the metabolite workflow.isn’t it?
As I said, it depends. If I were you, I would leave them out for now.
But some people like to align the blanks to remove features that also appear in blank samples.
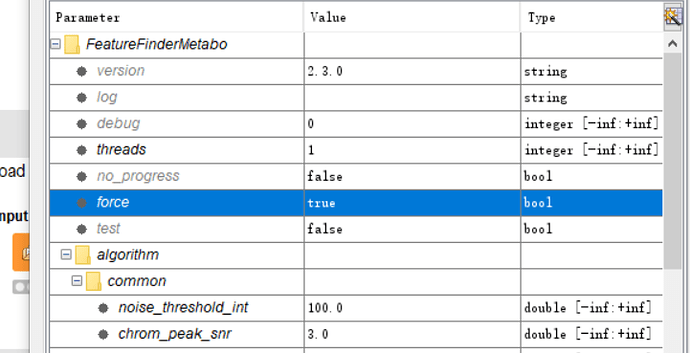
Now,I am testing the workflow with the data(A,B,C,D).The data have about 32 files,The workflow excutes slowly,I think it may be influenced by the many files. and I choose the parameters the force is true because it appears the data is profile data not centroid data.
Profile data makes it very slow and is not recommended. Instead of using “force” I would add a PeakPickerHiRes in front of your FeatureFinder, to centroid your data.
Yes,you are right. It excute slowly.But I converted the raw data into the data to centroid by the MSconvertGUI.In general, the data should be the centroid data by the MsconvertGUI.
Then I add the PeakPickerHiRes, it becomes quickly.
Weird, maybe it is a bug in MSConvertGui then? That it does not annotate correctly? I don’t know. To be safe, just keep the PeakPicker. It will ignore spectra that are already centroided.
Ok,I will keep the node.
This topic was automatically closed 90 days after the last reply. New replies are no longer allowed.