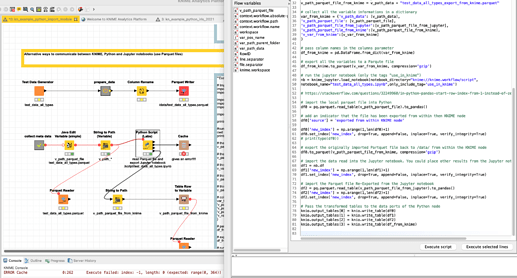
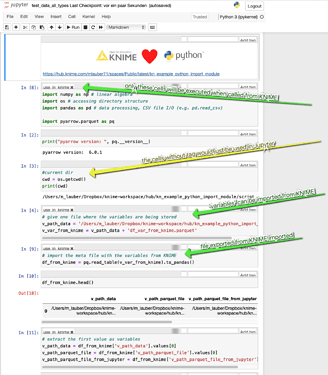
OK I have added a part to the workflow where KNIME, Python and Jupyter would communicate with each other. ‘Variables’ can be exchanged via a file and also data can be transferred via Parquet files. It would be great though to be able to transfer ‘real’ variables between KNIME and Jupyter (like in this idea).
Unfortunately on my Mac the export thru the new Python integration produces a strange error about index when trying to access the results. Some nodes (like Group by) although would work. I have tried several things with columnar backend, Flatbuffer communication and also re-setting the index in the pandas data frames. To no avail.
ERROR Cache 0:262 Execute failed: index: -1, length: 0 (expected: range(0, 364))
@MarcelW maybe you could investigate. My fear is that something with the arrow integration still is not right - admittedly I tried to challenge the parquet format somewhat with different data types (had to cut back on that already). But what is strange that the Parquet reader can handle that and also ‘within’ the Python node everything seems fine. What fails is the write_table:
knio.output_tables[1] = knio.write_table(df1)
But the ‘communication’ between KNIME, Python node in KNIME and Jupyter would just work fine.
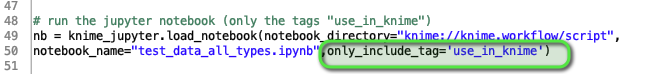
The interesting feature being to tag only the cells in the notebook “test_data_all_types.ipynb” in the subfolder “/script/” that you would want be executed in KNIME. So you could combine some exploration and execution.