A few things. The paths seem to be different on MacOS and Windows so you end up with a path with / and \ which is not good (maybe the os.sep is not adequate). And your pickle files have different names (did not immediately spot that).
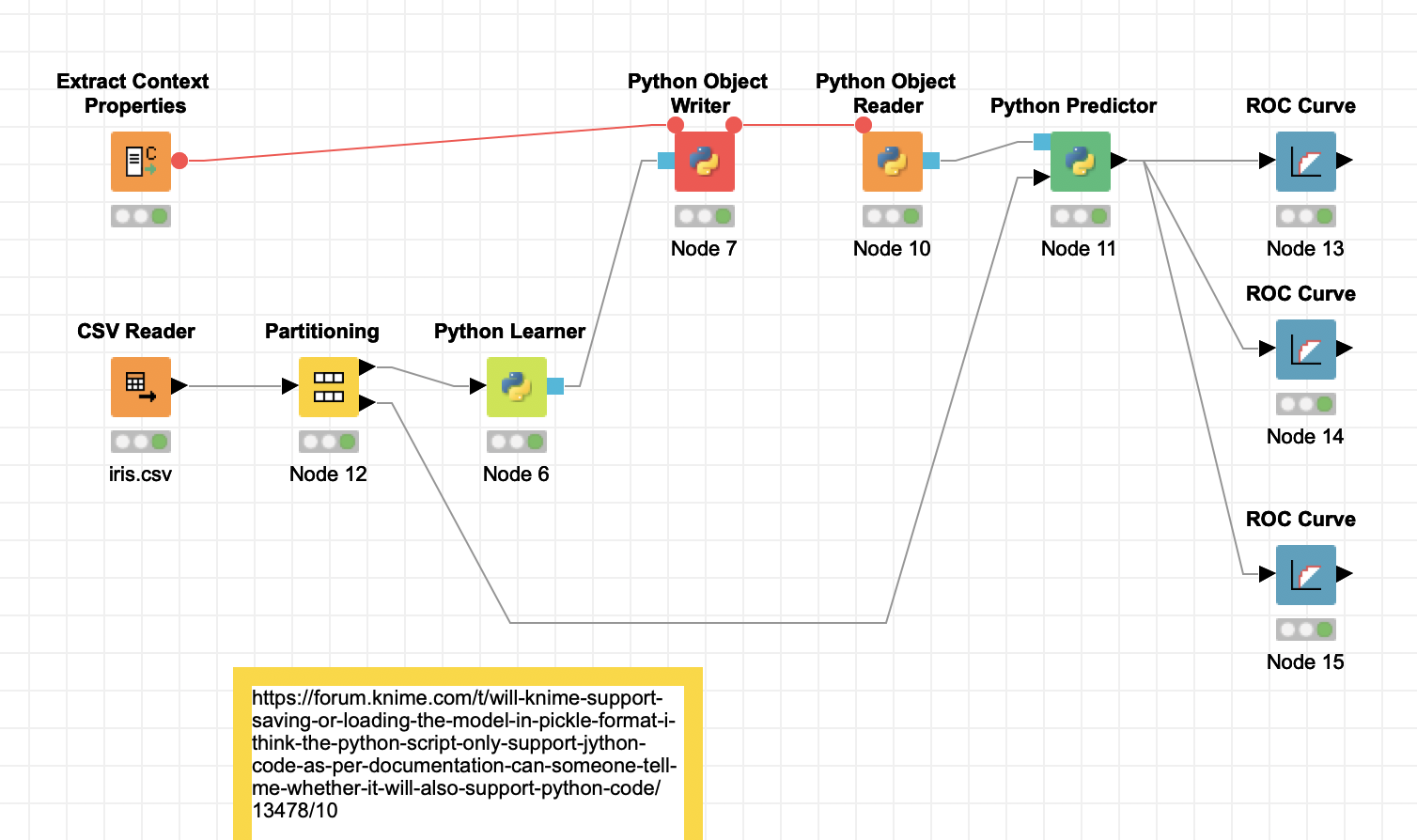
Attached a new modified workflow that does work.