Hello,
Does anyone have a workflow that they are willing to share that will append structural alerts for toxicity, PAINS, etc. to a table of input structures?
Thanks and regards,
Tim Ritchie.
Hi Tim,
There is a nice workflow on KNIME Public Examples server under 50_Applications/32_Hitlist_Processing/01_Hitlist_Filter
It uses REOS and PAINS filters.
You could use the RDKit Molecule Catalog Filter node and the RDKit Functional Group Filter node to search molecules using established sets or specify the fragments yourself
Best,
Daria
Hello Daria,
Many thanks for the suggestions.
I will take at look at the workflow. I wasn’t aware of the Functional Group Filter in RDKit.
Regards,
Tim.
Daria,
I received a sample workflow from Greg Landrum to essentially filter out PAINS compounds from an SD file that looks like this:
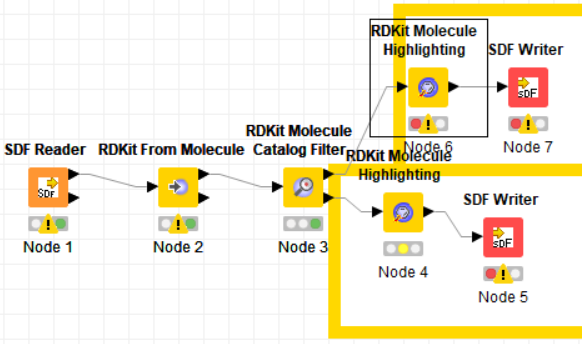
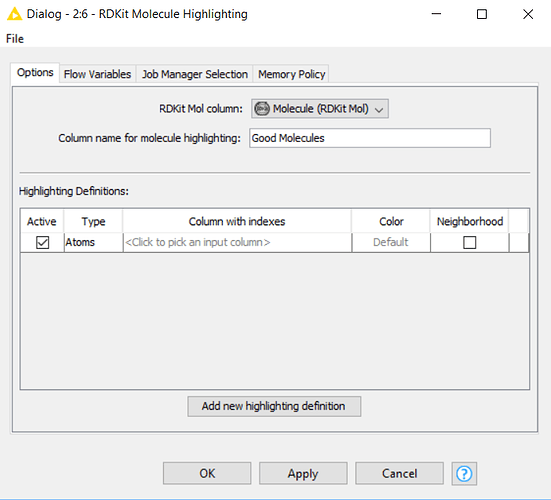
My problem is that while everything works fro the “bad molecules”. I am having problems executing the top “good molecules” workflow because of the highlighter node has no collection type column found in the input table which would contain atoms to be highlighted.
The column with indexes does not have any choices to choose from on the “good molecules” highlighter node.
Please advise…
Will