Dear Forum,
I am trying to handle rather large amounts of data from DNA sequencing. Or rather not the sequences themselves but some identifier text strings and associated counts, frequencies etc. I.e. few columns but many rows (100+ mio).
I can read in sub-pools of the files, concatenate them, but when I try to write the combined table as a csv, I observe a memory run-away and my server gulps up all the 56 GB RAM assigned to KNIME and the csv writer node fails (or KNIME freezes up). I have also tried to use a table writer node, but with same result. The 8 csv files that I’m trying to combine have a total size of ~3GB.
Any suggestions on how to do this or is there an upper limit somewhere? Appreciate you suggestions.
Best,
Kolster
Try to use this node before CSV Writer.
Hi there!
Sry for inconvenience.
Can you share a bit more info? What KNIME version are you using and what OS? When KNIME doesn’t freeze what error message do you get?
Is there possibility you can share workflow/data in order to test it?
Br,
Ivan
I have handled a table with around 325,000,000 rows which created a .table file on disk of about 1.6GB. The system used has 64GB RAM. Performance was not good. Reading or writing the table would go quickly for a while (in terms of the progress bar) and then stall. But it did eventually complete.
Hi there!
Sry to hear that @acommons but seems to me 325 million rows is a lot. Do you know approximatelly how long the writing lasted? Did you have a lot of columns? Regarding data types, were they standard like strings and numbers or some special types?
Br,
Ivan
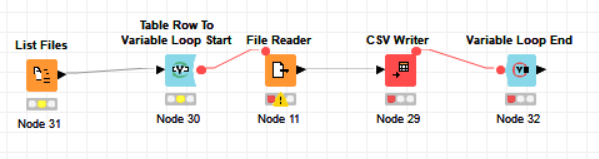
Could you try and write the data to CSV in loops? Only so much at a time and then append it to an existing CSV file. We had a similar discussion recently:
Hi there,
Sorry for not responding that quickly, but just wanted to let you know that writing larger csv files in chunks worked. I did set it up to read a series of files into a single one. Still puzzled why there seem to be a memory runaway when writing the larger files which were concatenated prior to csv writer node…
The only obstacle I encountered was to keep my column header and on the other hand not having headers in the middle of my combined file. Found a way around it by having a “header” csv with two rows of headers and a config file stating the other files to combine. Now set this up in a loop with a csv writer node in append mode
This topic was automatically closed 182 days after the last reply. New replies are no longer allowed.