Hello Knime users,
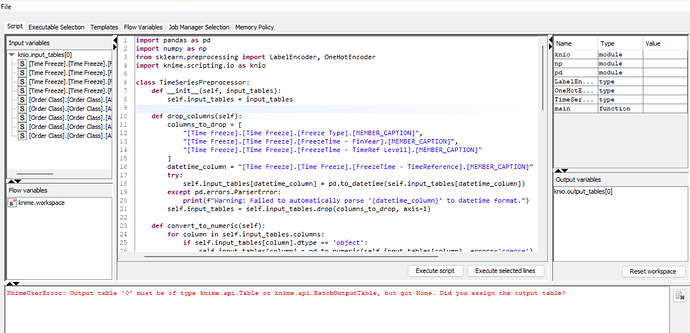
this is my preprocessing code. but after execute code in python script node of knime. then show error. please help me
KnimeUserError: Output table ‘0’ must be of type knime.api.Table or knime.api.BatchOutputTable, but got None. Did you assign the output table?
import pandas as pd
import numpy as np
from sklearn.preprocessing import LabelEncoder, OneHotEncoder
import knime.scripting.io as knio
class TimeSeriesPreprocessor:
def init(self, input_tables):
self.input_tables = input_tables
def drop_columns(self):
columns_to_drop = [
"[Time Freeze].[Time Freeze].[Freeze Type].[MEMBER_CAPTION]",
"[Time Freeze].[Time Freeze].[FreezeTime - FinYear].[MEMBER_CAPTION]",
"[Time Freeze].[Time Freeze].[FreezeTime - TimeRef Level1].[MEMBER_CAPTION]"
]
datetime_column = "[Time Freeze].[Time Freeze].[FreezeTime - TimeReference].[MEMBER_CAPTION]"
try:
self.input_tables[datetime_column] = pd.to_datetime(self.input_tables[datetime_column])
except pd.errors.ParserError:
print(f"Warning: Failed to automatically parse '{datetime_column}' to datetime format.")
self.input_tables = self.input_tables.drop(columns=columns_to_drop, axis=1) # Corrected the drop method
def convert_to_numeric(self):
for column in self.input_tables.columns:
if self.input_tables[column].dtype == 'object' and column != "[Time Freeze].[Time Freeze].[FreezeTime - TimeReference].[MEMBER_CAPTION]":
self.input_tables[column] = pd.to_numeric(self.input_tables[column], errors='coerce')
def convert_numeric_objects(self):
for column in self.input_tables.columns:
if self.input_tables[column].dtype == 'object' and column != "[Time Freeze].[Time Freeze].[FreezeTime - TimeReference].[MEMBER_CAPTION]":
self.input_tables[column] = self.input_tables[column].astype(float)
def encode_categorical(self):
for column in self.input_tables.columns:
if self.input_tables[column].dtype == 'object' and column != "[Time Freeze].[Time Freeze].[FreezeTime - TimeReference].[MEMBER_CAPTION]":
if len(self.input_tables[column].unique()) <= 10:
le = LabelEncoder()
self.input_tables[column] = le.fit_transform(self.input_tables[column])
else:
ohe = OneHotEncoder(handle_unknown='ignore', sparse=False)
encoded_data = pd.DataFrame(ohe.fit_transform(self.input_tables[[column]]))
self.input_tables = pd.concat([self.input_tables, encoded_data], axis=1)
self.input_tables.drop(column, axis=1, inplace=True)
def fill_missing_values(self):
self.input_tables = self.input_tables.interpolate(method='linear')
def handle_outliers(self):
for column in self.input_tables.columns:
if self.input_tables[column].dtype != 'datetime64[ns]' and column != "[Time Freeze].[Time Freeze].[FreezeTime - TimeReference].[MEMBER_CAPTION]":
q1 = self.input_tables[column].quantile(0.25)
q3 = self.input_tables[column].quantile(0.75)
iqr = q3 - q1
lower_bound = q1 - 1.5 * iqr
upper_bound = q3 + 1.5 * iqr
self.input_tables[column] = np.where(self.input_tables[column] < lower_bound, lower_bound, self.input_tables[column])
self.input_tables[column] = np.where(self.input_tables[column] > upper_bound, upper_bound, self.input_tables[column])
def preprocess_data(self):
self.drop_columns()
self.convert_to_numeric()
self.convert_numeric_objects()
self.encode_categorical()
self.fill_missing_values()
self.handle_outliers()
return self.input_tables # Return the preprocessed data
def main():
input_tables = knio.input_tables[0].to_pandas()
preprocessor = TimeSeriesPreprocessor(input_tables)
preprocessed_data = preprocessor.preprocess_data()
output_tables = knio.Table.from_pandas(preprocessed_data)
knio.output_tables[0] = output_tables
if name == “main”:
main()