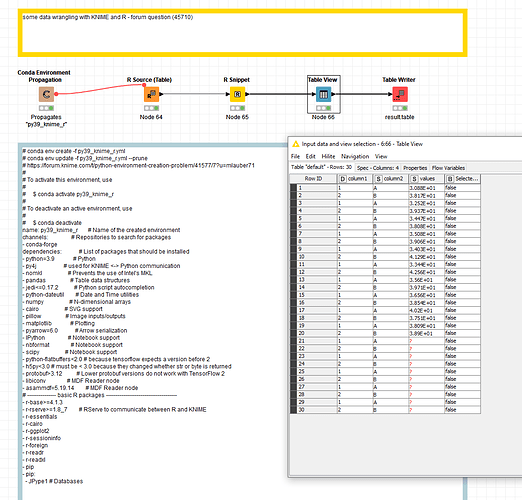
@cuongnguyen maybe you try again with some more basic settings and a newer R version (4.1.3) which is available via conda(-forge). Your environment propagation has a lot of special linux packages that I cannot install on Windows/MacOS - so I set up a more generic YAML file - you could add other packages and update the environment later.
The YML/YAML file would look like this
# conda env create -f py39_knime_r.yml
# conda env update -f py39_knime_r.yml --prune
# https://forum.knime.com/t/python-environment-creation-problem/41577/7?u=mlauber71
# https://docs.knime.com/latest/r_installation_guide/index.html
#
# To activate this environment, use
#
# $ conda activate py39_knime_r
#
# To deactivate an active environment, use
#
# $ conda deactivate
name: py39_knime_r # Name of the created environment
channels: # Repositories to search for packages
- conda-forge
dependencies: # List of packages that should be installed
- python=3.9 # Python
- py4j # used for KNIME <-> Python communication
- nomkl # Prevents the use of Intel's MKL
- pandas # Table data structures
- jedi<=0.17.2 # Python script autocompletion
- python-dateutil # Date and Time utilities
- numpy # N-dimensional arrays
- cairo # SVG support
- pillow # Image inputs/outputs
- matplotlib # Plotting
- pyarrow=6.0 # Arrow serialization
- IPython # Notebook support
- nbformat # Notebook support
- scipy # Notebook support
- python-flatbuffers<2.0 # because tensorflow expects a version before 2
- h5py<3.0 # must be < 3.0 because they changed whether str or byte is returned
- protobuf>3.12 # Lower protobuf versions do not work with TensorFlow 2
- libiconv # MDF Reader node
- asammdf=5.19.14 # MDF Reader node
# --------------- basic R packages -------------------------------------
- r-base>=4.1.3
- r-rserve>=1.8_7 # RServe to communicate between R and KNIME
- r-essentials
- r-cairo
- r-ggplot2
- r-sessioninfo
- r-foreign
- r-readr
- r-readxl
- pip
- pip:
- JPype1 # Databases