Dear all,
i was wondering how to perform in knime some manipulations on molecular structures, for example pruning exocyclic double bonds (including carbonyl groups) and substituents (often called also R groups).
Not sure if a RDkit one component reaction node would suit my needs, as i wasn’t not able to set up a reasonable SMARTS query.
Thank you in advance
Do you have specific examples of structures you want to modify and the specific modifications you’d like to carry out?
@C_C_85 Have a look at the discussion in this thread for some guidance:
(the other)
Simon
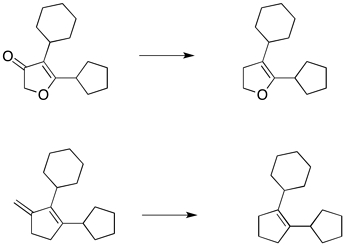
Basically, for a list of different molecules, i need to remove side chains and terminal atoms from rings. This pruning procedure would give for each molecule a more generalised substructure. I attached an example here.
Do all of your molecules share the same core/scaffold? If not, how may cores/scaffolds are there?
Have you tried using any of the R-Group Decomposition nodes or MCS nodes?
I have a list of molecules bearing different cores, but still, among them there are small series sharing the same core. The R-Group Decomposition might actually do what i need, but still i can’t figure out how to remove the R-groups from the molecule. The output gives a Core with “A” replacing the R groups; i tried the Feature remover to try removing pseudoatoms or attachment points, but it gives me a NullPointerException
I find the R-Group Decomposition node to be a little finicky.
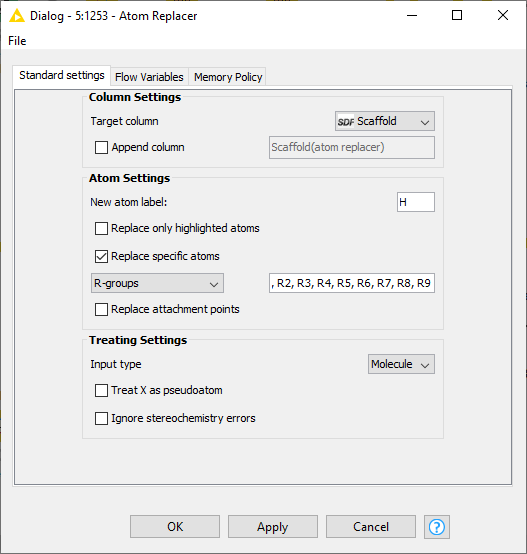
I prefer the R-Group Decomposer node. It replaces the attachment points with numbered R groups which I can replace using the Atom Replacer node:
Very interesting solution,
i wish i could see the result but i’m struggling with errors raised by the Decomposer node (and in general with Indigo nodes), maybe it happens because of the molecule type cast nodes upstream?!
Can you share your workflow here?
Yes,
the protocol attachedR-group_decomposition_protocol.knwf (35.4 KB) contains just few nodes, the SMILES string in the table creator is related to the example molecule of the image above.
Besides the substituents, I’d like to remove also the carbonyl group from the ring.
I’m using knime 4.3.2.
Thank you!
This is a difficult problem. I can strip down the core to the most basic version, which includes only single bonds and a carbon atoms framework, but I haven’t found a way to preserve the double bond or the oxygen atom like in the picture you posted earlier.
True, i’m afraid that it can’t be accomplished with the available nodes…