We have just released another update to the nightly build.
In this latest update, we have retired the existing PMI Calculation node which has been in the public domain since v1.6.4 (April 2016), and replaced it with a new node Principal Moment of Intertia (PMI)-Derived Properties. This uses a new underlying framework, and is no longer dependant on RDKit. It also now accepts CTab and PDB cells as inputs. The new framework also sets column properties with a brief description, units, literature references, aliases and minimum and maximum values where available. The node also calculates many more PMI-related properties (20 in total). The properties are:
- PMI 1 - First (smallest) Principal Moment of Inertia (also referred to as: I₁)
- PMI 2 - Second Principal Moment of Inertia (also referred to as: I₂)
- PMI 3 - Third (largest) Principal Moment of Inertia (also referred to as: I₃)
- npr1 - First Normalised PMI (i.e. I₁ / I₃) (See: Sauer and Schwartz, J. Chem. Inf. Comput. Sci., 2003, 43, 987-1003)
- npr2 - Second Normalised PMI (i.e. I₂ / I₃) (See: Sauer and Schwartz, J. Chem. Inf. Comput. Sci., 2003, 43, 987-1003)
- Sigma npr - npr1 + npr2 (also referred to as: Flatsum, npr1+npr2, Σnpr) (See: )
- Sphericity - npr1 + npr2 - 1 (also referred to as: S) (See: Wirth et al., J. Comput. Aided Mol. Des., 2013, 27, 511-524)
- Rodlikeness - npr2 - npr1 (also referred to as: R, Rod-likeness) (See: Wirth et al., J. Comput. Aided Mol. Des., 2013, 27, 511-524)
- Disklikeness - 2 - 2 × npr2 (also referred to as: D, Disc-likeness) (See: Wirth et al., J. Comput. Aided Mol. Des., 2013, 27, 511-524)
- Asphericity - 0.5 × [(I₁ - I₂)² + (I₁ - I₃)² + (I₂ - I₃)²]/[I₁² + I₂² + I₃²]; 0 corresponds to a spherical top and 1 to a linear molecule. ~0.25 corresponds to prolate (cigar-shaped) molecules, and disc-shaped ~1 (also referred to as: ΩA) (See: Arteca, Reviews in Computational Chemistry, Vol 9 VCH, NY, 1991, 191-253)
- Inertial Shape Factor - I₂ / (I₁ × I₃); Undefined for planar molecules (also referred to as: SI) (See: Lister et al., Internal Rotation and Inversion, Academic Press, London, 1978)
- Molecule Eccentricity - √(I₁² - I₃²) / I₂; 0 corresponds to a spherical top, and 1 to a linear molecule (also referred to as: ε) (See: Arteca, Reviews in Computational Chemistry, Vol 9 VCH, NY, 1991, 191-253)
- Gyradius - Radius of Gyration (also referred to as: RG, Radius of Gyration, Gyradius) (See: )
- Gyradius 2D - Radius of Gyration (2D) = √(I₁ × I₂) / MWt (also referred to as: RG, Radius of Gyration, Gyradius) (See: Volkenstein ‘Configurational Statistics of Polymeric chains’ Wiley, NY, 1963)
- Gyradius 3D - Radius of Gyration (3D) = ∛(I₁ × I₂ × I₃) / MWt (also referred to as: RG, Radius of Gyration, Gyradius) (See: Tanford ‘Physical Chemistry of Macromolecules’, Wiley, NY, 1961)
- PRG 0 - 1st Principle Radius of Gyration (R0 = √(I₁ / MWt)) (also referred to as: R0) (See: Jian et al., ‘A dimension map for molecular aggregates’, J. Mol. Graphics. Model., 2015, 58, 10-15)
- PRG 1 - 2nd Principle Radius of Gyration (R1 = √(I₂ / MWt)) (also referred to as: R1) (See: Jian et al., ‘A dimension map for molecular aggregates’, J. Mol. Graphics. Model., 2015, 58, 10-15)
- PRG 2 - 3rd Principle Radius of Gyration (R2 = √(I₃ / MWt)) (also referred to as: R2) (See: Jian et al., ‘A dimension map for molecular aggregates’, J. Mol. Graphics. Model., 2015, 58, 10-15)
- Gyradius Ratio 1 - 1st Gyradius Ratio (r1 = PRG1 / PRG0) (also referred to as: r1) (See: Jian et al., ‘A dimension map for molecular aggregates’, J. Mol. Graphics. Model., 2015, 58, 10-15)
- Gyradius Ratio 2 - 2nd Gyradius Ratio (r2 = PRG2 / PRG0) (also referred to as: r2) (See: Jian et al., ‘A dimension map for molecular aggregates’, J. Mol. Graphics. Model., 2015, 58, 10-15)
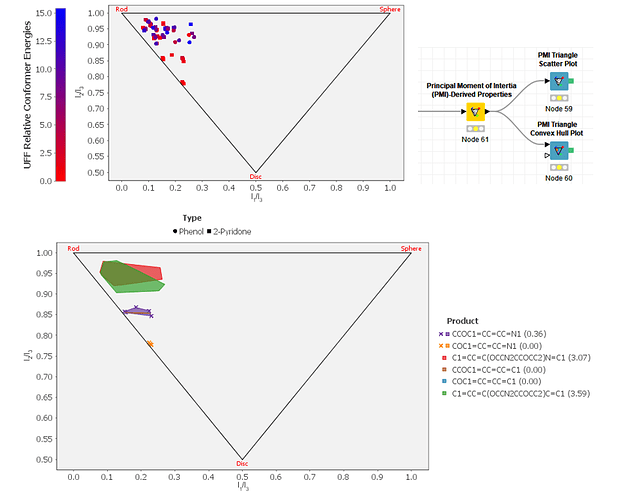
We have also added the first two of a family of PMI Plotting nodes, based on the JFreeChart integration withing KNIME. These first two nodes provide a standard PMI Scatter plot, and a Convex Hull PMI Plot (which can have an optional scatter point set overlaid onto it):
There will be more releases with further new nodes in the coming days.
Yours,
Steve